Gruiformes (rails, cranes & allies)
The taxon comprises ~192 species in ~6 families.

Genus-level timetree of extant Gruiformes according to Prum et al. (2015), Boast et al. (2019), Garcia-R. et al. (2020), Garcia-R. & Matzke (2021), Kuhl et al. (2021), Kirchman et al. (2021),
Depino et al. (2023), and Stiller et al. (2024), with the
distribution of each taxon being indicated by the colour-code used throughout this website (Distribution
code). Primary
divergence times follow Stiller et al. (2024): (Heliornis + (Atlantisia/Laterallus + Zapornia)) + (Psophia + (Aramus + (Balearica + Grus))).
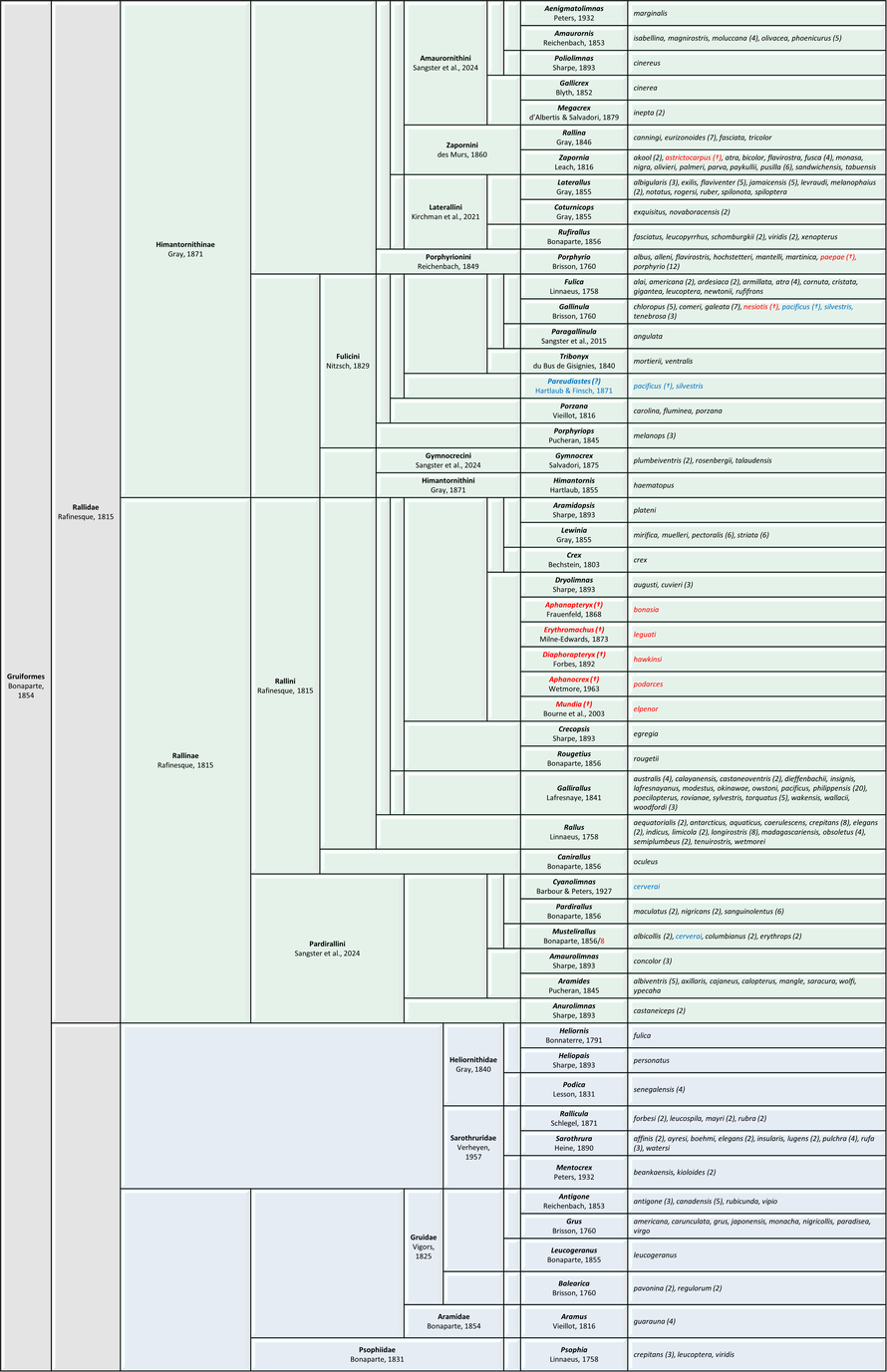
Traditional genus-level classification of extant Gruiformes following the AviList checklist v2025. (link) The number of subspecies is given in parentheses.
References
Akiyama T, Nishida C, Momose K, Onuma M, Takami K, and Masuda R (2017), Gene duplication and concerted evolution of mitochondrial DNA in crane species, Mol. Phylogenet. Evol. 106, 158-163. (abstract)
Boast AP, Chapman B, Herrera MB, Worthy TH, Scofield RP, Tennyson AJD, Houde P, Bunce M, Cooper A, and Mitchell KJ (2019), Mitochondrial genomes from New Zealand´s extinct adzebills (Aves: Aptornithidae: Aptornis) support a sister-taxon relationship with the Afro-Madagascan Sarothruridae, Diversity 11, e:24. (free pdf)
Chaves JA, Martinez-Torres PJ, Depino EA, Espinoza-Ulloa S, García-Loor J, Beichman AC, and Stervander M (2020), Evolutionary history of the Galápagos Rail revealed by ancient mitogenomes and modern samples, Diversity 12, e:425. (free pdf)
Chen P, Huang Z, Zhu C, Han Y, Xu Z, Sun G, Zhang Z, Zhao D, Ge G, and Ruan L (2020), Complete mitochondrial genome and phylogenetic analysis of Gruiformes and Charadriiformes, Pakistan J. Zool. 52, 425-439. (free pdf)
Depino EA, Pérez-Emán JL, Bonaccorso E, and Areta JI (2023), Evolutionary history of New World crakes (Aves: Rallidae) with emphasis on the tribe Laterallini, Zool. Scr. 52, 394-412. (abstract)
Garcia-R. JC, Gibb GC, and Trewick SA (2014), Deep global evolutionary radiation in birds: diversification and trait evolution in the cosmopolitan bird family Rallidae, Mol. Phylogenet. Evol. 81, 96-108. (abstract)
Garcia-R. JC, and Trewick SA (2015), Dispersal and speciation in purple swamphens (Rallidae: Porphyrio), Auk 132, 140-155. (free pdf)
Garcia-R. JC, Lemmon EM, Lemmon AR, and French N (2020), Phylogenomic reconstruction sheds light on new relationships and timescale of rails (Aves: Rallidae) phylogeny, Diversity 12, e:70. (pdf)
Garcia-R. JC, and Matzke NJ (2021), Trait-dependent dispersal in rails (Aves: Rallidae): historical biogeography of a cosmopolitan bird clade, Mol. Phylogenet. Evol. 159, e:107106. (abstract)
Groenenberg DSJ, Beintema AJ, Dekker RWRJ, and Gittenberger E (2008), Ancient DNA elucidates the controversy about the flightless island hens (Gallinula sp.) of Tristan da Cunha, PLoS ONE 3, e:1835. (pdf)
Kirchman JJ (2012), Speciation of flightless rails on islands: a DNA-based phylogeny of the typical rails of the Pacific, Auk 129, 56-69. (free pdf)
Kirchman JJ, Rotzel McInerney N, Giarla TC, Olson SL, Slikas E, and Fleischer RC (2021), Phylogeny based on ultra-conserved elements clarifies the evolution of rails and allies (Ralloidea) and is the basis for a revised classification, Ornithology, 2021, e:ukab041. (free pdf)
Krajewski C, Sipiorski JT, and Anderson FE (2010), Complete mitochondrial genome sequences and the phylogeny of cranes (Gruiformes: Gruidae), Auk 127, 440-452. (free pdf)
Krajewski
C (2019),
Phylogenetic taxonomy of cranes and the evolutionary origin of the whooping crane, In: Nyhus PJ, French JB, Converse SJ, Austin JE, and Delap JH (eds), “Whooping cranes: biology and conservation.
Biodiversity of the world: conservation from genes to landscapes”, p. 17-24. Academic Press, Cambridge. (link)
Kuhl H, Frankl-Vilches C, Bakker A, Mayr G, Nikolaus G, Boerno ST, Klages S, Timmermann B, and Gahr M (2021), An unbiased molecular approach using 3'UTRs resolves the avian family-level tree of life, Mol. Biol. Evol. 38, 108-127. (free pdf)
Lian T, Yang C, Yuan H, Wang QX, Du XJ, and Li XJ (2021), Characterization of the complete mitogenomes of Baillon‘s Crake Porzana pusilla and phylogenetic analysis, Mitochondrial DNA Part B 6:2. (pdf)
Maley JM, and Brumfield RT (2013), Mitochondrial and next-generation sequence data used to infer phylogenetic relationships and species limits in the Clapper/King Rail complex, Condor 115, 316-329. (free pdf)
Mayr G (2019), Hypotarsus morphology of the Ralloidea supports a clade comprising Sarothrura and Mentocrex to the exclusion of Canirallus, Acta Ornithol. 54, 51-58. (abstract)
Musser GM, and Cracraft J (2019), A new morphological dataset reveals a novel relationship for the adzebills of New Zealand (Aptornis) and provides a foundation for total evidence neoavian phylogenetics, Am. Mus. Novit. 3927, 1-70. (free pdf)
Musser G, Ksepka DT, and Field DJ (2019), New material of Paleocene-Eocene Pellornis (Aves: Gruiformes) clarifies the pattern and timing of the extant gruiform radiation, Diversity 11, e:102. (free pdf)
Nováková N, and Robovsky J (2021), Behaviour of cranes (family Gruidae) mirrors their phylogenetic relationships, Avian Res. 12: e:40. (pdf)
Oswald JA, Terrill RS, Stucky BJ, LeFebvre MJ, Steadman DW, Guralnick RP, and Allen JM (2021), Ancient DNA from the extinct Haitan cave-rail (Nesotrochis staganinos) suggests a biogeographic connection between the Caribbean and Old World, Biol. Lett. 17, e:20200760. (pdf)
Prum RO, Berv JS, Dornburg A, Field DJ, Townsend JP, Lemmon EM, and Lemmon AR (2015), A comprehensive phylogeny of birds (Aves) using targeted next-generation DNA sequencing, Nature 526, 569-57. (abstract)
Sangster G, Garcia-R. JC, and Trewick SA (2015), A new genus for the Lesser Moorhen Gallinula angulata Sundevall, 1850 (Aves, Rallidae), Eur. J. Taxon. 153, 1-8. (pdf)
Sangster G, Blokland JC, Gregory SMS, and Dickinson EC (2024), Gymnocrecini, Amaurornithini and Pardirallini: three new family-group names for rails, with comments on the taxonomic placement of Zapornia akool (Rallidae), Avian Syst. 2, 53-64. (pdf)
Slikas B, Olson SL, and Fleischer RC (2002), Rapid, independent evolution of flightlessness in four species of Pacific Island rails (Rallidae): an analysis based on mitochondrial sequence data, J. Avian Biol. 33, 5-14. (free reading)
Stervander M, Ryan PG, Melo M, and Hansson B (2019), The origin of the world´s smallest flightless bird, the Inaccessible Island Rail Atlantisia rogersi (Aves: Rallidae), Mol. Phylogenet. Evol. 130, 92-98. (abstract)
Stiller J, Feng S, Chowdhury AA, Rivas-González I, Duchêne DA, Fang Q, Deng Y, Kozlov A, Stamatakis A, Claramunt S, Nguyen JMT, Ho SYW, Faircloth BC, Haag J, Houde P, Cracraft J, Balaban M, Mai U, Chen G, Gao R, Zhou C, Xie Y, Huang Z, Cao Z, Yan Z, Ogilvie HA, Nakhleh L, Lindow B, Morel B, Fjeldså J, Hosner PA, da Fonseca RR, Petersen B, Tobias JA, Székely T, Kennedy JD, Reeve AH, Liker A, Stervander M, Antunes A, Tietze DT, Bertelsen M, Lei F, Rahbek C, Graves GR, Schierup MH, Warnow T, Braun EL, Gilbert MTP, Jarvis ED, Mirarab S, and Zhang G (2024), Complexity of avian evolution revealed by family-level genomes, Nature 629, 851-860. (pdf)
Verry AJF, Mas-Carrió E, Gibb GC, Dutoit L, Robertson BC, Waters JM, and Rawlence NJ (2024), Ancient mitochondrial genomes unveil the origins and evolutionary history of New Zealand‘s enigmatic takahe and moho, Mol. Ecol. 33, e:17227. (pdf)
Wang T, Wang H, Zhao Z, Wang Z, Mu L, and Yu H (2018), Complete mitochondrial genome of the Siberian Crane Grus leucogeranus, Mitochondrial DNA Part B 3, 575-576. (pdf)
Yang R, Wu X, Yan P, Su X, and Yang B (2009) Complete mitochondrial genome of Otis tarda (Gruiformes: Otididae) and phylogeny of Gruiformes inferred from mitochondrial DNA sequences, Mol. Biol. Rep. 37, 3057-66. (abstract)

